A whole new class of molecules make conventional protein antibodies obsolete but are they safe? Dr Mae-Wan Ho
Aptamers are making conventional antibodies obsolete. Aptamers are oligonucleic acids or peptides that bind to a specific target molecule like conventional protein antibodies but with little or no immunogenicity and improved stability, and much easier and cheaper to produce in the test-tube, without the need for cells or animals [1].
Nucleic acid aptamers, in particular, are finding numerous applications that previously required the use of antibodies. They are engineered through repeated rounds of in vitro selection or SELEX (see Box 1) to bind to various targets such as small molecules, proteins, nucleic acids, even cells, tissues, and organisms.
Box 1
SELEX, an in vitro selection technique for producing aptamers [2]
SELEX (systematic evolution of ligands by exponential enrichment) is a method for producing oligonucleotides of single stranded DNA or RNA that specifically bind to a target ligand. It begins with the synthesis of a very large oligonucleotide library consisting of randomly generated sequences of fixed lengths. For length n, the number of possible sequences in the library is n4, because there are four different bases possible at each position. The sequences in the library are exposed to the target, which may be a protein, or a small organic compound. Those that do not bind are removed. The bound sequences are recovered and amplified by PCR to prepare for another round of selection in which the stringency of binding conditions is increased, and so on.
Aptamers were first produced independently in two laboratories in 1990, that of Larry Gold [3] at University of Colorado, Boulder, who used the term SELEX (systematic evolution of ligands by exponential enrichment) for the process of selecting RNAs binding to T4 DNA polymerase, and Jack Szostak [4] at Massachusetts General Hospital, Boston, selecting for RNAs binding to various organic dyes, and coined the terms ‘in vitro selection’ and ‘aptamer’.
The technique has been used to evolve oligonucleotide aptamers of extremely high binding affinity to a variety of targets, such as ATP and adenosine, and proteins such as prion and vascular endothelial growth factor (VEGF). Clinical uses are suggested for aptamers that bind tumour markers, and a VEGF-binding aptamer trade-named Macugen has been approved by the FDA for treating macular degeneration (see below).
Aptamers bind to ligands based on their three-dimensional structure; hence different oligonucleotide sequences may recognize the same target molecules.
However, selection for extremely high binding affinities (at 10-9M or less) does not guarantee absolute specificity. Off-target binding may have significant, unintended clinical effects [5]. This is particularly important as natural aptamers have been discovered that play key roles in gene regulation.
Natural aptamers were discovered in 2002 in the nucleic acid genetic regulatory element, the riboswitch that possesses similar molecular recognition properties to the artificially made aptamers [6]. A riboswitch is a part of an mRNA molecule – transcribed from a gene -that directly binds to a small target molecule, usually a metabolite, to change the gene’s activity [7]. This involves turning off gene expression via the premature termination of transcription, or inhibition of translation; or the riboswitch may be an RNA enzyme (ribozyme) that cleaves itself in the presence of the small metabolite, or it may lead to alternative splicing of the pre-mRNA. Some riboswitches may even regulate the transcription of downstream genes; or turn on gene expression as they bind to target molecules.
Many of the earliest riboswitches corresponded to conserved sequence motifs in the 5’ untranslated regions of mRNAs. Most known riboswitches occur in viruses and eubacteria, but they have also been discovered in plants and certain fungi, and predicted in archaea bacteria. The first human riboswitch was identified in 2009 in human vascular endothelial growth factor (VEGF) [8]. It is highly likely that riboswitches may also exist in other animals.
The many uses for aptamers are summarized in Figure 1 [9].
Figure 1 Biological applications of aptamers
Oligonucleic aptamers are replacing conventional protein antibodies because they are much more stable, easy and cheap to produce, less immunogenic or toxic and can target a wider range of molecules, including those not giving strong conventional antibody responses.
Protein antibodies are easily denatured and lose their structure irreversibly at high temperatures. Oligonucleotide aptamers in contrast are much more heat- stable, and maintain their structures over repeated cycles of denaturation/renaturation. Aptamers can therefore be used under a wider range of assay conditions.
Monoclonal antibodies are laborious and expensive to produce, requiring large scale mammalian cell cultures; and different batches have to be tested to ensure that they maintain the same binding properties. In contrast, aptamers, once selected, can be synthesized in bulk cheaply with accuracy and reproducibility by chemical reactions. Aptamers can also be readily modified to increase their stability and resistance to nuclease breakdown (though this runs the danger that they will persist in the body and in the environment to do harm), and it is possible to introduce signalling adducts such as fluorescent molecule and its quencher for biosensing.
Protein antibodies are immunogenic, which makes repeated dosing a big problem. Aptamers are reported to be low in immunogenicity and toxicity, and are readily broken down in the body, unless modified to prevent breakdown. However, the evidence for low immunogenicity and toxicity appears to be based solely on the Phase I and Phase II clinical trials of mucagen [9, 10], which may well not be generalizable to the entire class of molecules.
Finally, it is often difficult to produce antibodies against small molecules, ions, and toxins that do not elicit strong immune response; but aptamers can be generated that bind to those molecules with high affinity. Aptamers not only substitute for conventional antibodies, but also have an almost unlimited potential to overcome the limitations of antibodies.
Biosensors based on aptamers as recognition elements are called aptosensors. Aptosensors can be made through a variety of techniques; based on changes in electrochemical potential, fluorescence (see Figure 2), or colour when the aptamer binds to the target molecule [9]. These aptosensors have found important applications in food safety, in the rapid detection of toxins and contaminants, such as antibiotics, pesticides, mycotoxins, heavy metals, bisphenol A, and adulterants such as illegal food dyes [11].
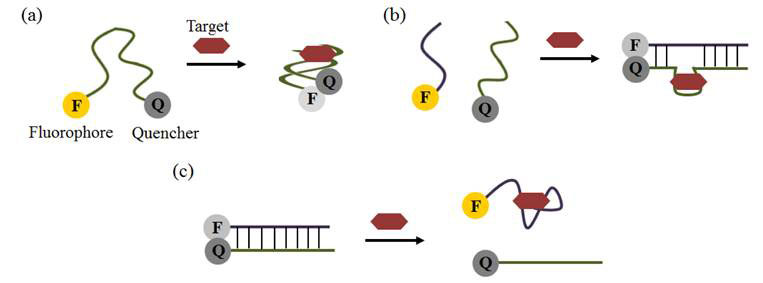
Figure 2 Aptosensors based on change in fluorescence: (a) fluorescence is quenched (decreased) when aptamer binds target, bringing fluorophore and quencher together; (b) aptamer binding causes complementary strands to pair up, again quenching fluorescence; (c) target binding separates complementary strands causing an increase in fluorescence
Aptosensors can also be used in clinical diagnosis in place of ELISA (enzyme-linked immunoadsorbent assay). In the Aptosensor version (ALISA), the aptamer is linked to biotin, which, on binding to the target immobilized on a nitrocellulose filter, reacts with streptavidin and horse radish peroxidase to generate the colour reaction. This and related techniques are offering commercially available rapid and simple methods for diagnosis of infectious diseases such as malaria and influenza [9].
Aptosensor microarrays have been designed that are particularly good for detecting cell and tissue markers for bio-imaging and drug delivery. Special light sensitive adduct enable aptamers to covalently link to bound proteins, making it easy to identify them [12].
Aptamers that bind to internalized cell surface receptors can deliver drugs and other cargoes into cells. This is very useful in cancer chemotherapy, which aims to kill cancer cells with cytotoxic drugs without harming noncancerous cells.
The prostate-specific membrane antigen PSMA is an important marker for prostate cancer. An aptamer combination for PSMA has been used to deliver doxorubicin, an anticancer drug into prostate cancer cells [13]. The anticancer drug has also been successfully delivered into liver- [14] and breast-cancer cells [15] by aptamers that target specific membrane antigens that are over-expressed in the respective cancer cells. A novel MUC1 aptamer selectively delivers the cytotoxic agent into both lung and breast cancer cells [16]; the MUC1 protein is over-expressed in most adenocarcinomas.
Aptamers are poised to make a big difference to cancer chemotherapy, which has been notorious for its toxic side effects on account of the difficulty in selectively targeting cancer cells.
Macugen, developed by Pfizer and Eyetech, is already commercially available for treating age-related macula degeneration, an eye disease that can lead to blindness. The drug is a polyethylene glycol-aptamer with specificity for VEGF165, which plays a critical role in angiogenesis (development of blood vessels) and permeability [17]. Regado Bioscience has developed a new aptamer drug for anticoagulation currently in Phase II clinical trials. The drug REG1 consists of two components: RB006 (coagulation factor IXa-specific aptamer) and RB007 (oligonucleotide antidote of the RB006 aptamer), to control excessive bleeding due to anticoagulation [18]. In the treatment of acute ischemic stroke (arising from a decrease in blood supply to the brain), a Factor IXa aptamer was found to improve neurological function in a mouse model, and treatment with its specific antidote in cases of intracranial haemorrhage improved survival [19]. These results highlight the need for antidote molecules in a growing number of therapeutic applications involving aptamers, which may well have off-target effects. Many are currently undergoing clinical trials, including a nuclein-specific aptamer for acute myeloid leukemia, a vonWillebrand factor-specific aptamer for carotid artery disease, and a thrombin-specific aptamer for anticoagulation [20, 21].
The potential number of aptamers for therapeutics is unlimited. A number of them target coagulation, and represent potential anticoagulants. But anticoagulants can cause complications such as considerable bleeding. Currently only an anticoagulant and antidote pair, heparin and protamine, is routinely used in clinics. This is now replaced by an anticoagulant aptamer and its antisense antidote.
Bruce Sullenger and colleagues at Duke University, Durham, North Carolina realized that with an ever increasing number of people taking numerous drugs, the need to safely administer drugs and limit unintended side effects has never been greater. Between 1998 and 2005, the number of serious adverse drugs events reported to the USFDA increased 2.6 fold and fatal adverse events increased 2.7-fold to 15 107 events in 2005.
Antidote control is the most direct way to counteract acute side effects of drugs, but it has been difficult and cost prohibitive to generate antidotes for most therapeutic agents. The team embarked on the development of a set of antidote molecules that are capable of counteracting the side effects of an entire class of therapeutics – aptamers – making them a particular safe class of therapeutics.
Previously, the team used Watson-Crick base-pairing rules to create a customized antidote oligonucleotide for each aptamer, which is very costly, and also results in a double-stranded RNA molecule that may stimulate the immune system. Instead, molecules that can sequester oligonucleotides in a sequence-independent manner should be able to function as universal antidotes for extracellular oligonucleotide-based drugs.
They screened a number of nucleic acid binding polymers for their ability to act as antidotes for a series of anticoagulant aptamers with very different structures. b-cyclodextrin-containing polycation (CDP) and polyphosphoramidate polymer (PPA-DPA) were found to act as sequence-independent antidotes for aptamers both in vitro and in vivo, in mice, and in pigs for CDP, without toxicity, and at lower level of 2.5mg/kg, compared with the conventional antidote protamine at 10mg/kg.
Nucleic acid aptamers appear to offer infinite opportunities for diagnosis, biosensing, bio-imaging, drug delivery, therapy, and much more. But key questions on safety remain largely unanswered. Although unmodified nucleic acid aptamers seem to be non-immunogenic and non-toxic, and rapidly degraded; the current trend is to chemically modify them to resist breakdown, hence increasing their therapeutic efficacy. But this also means that they are more likely to survive in the environment with unknown consequences for health and the ecology of our aquatic systems. This is of particular concern, as natural aptamers have been identified in numerous species including humans, and are very likely to be widely distributed in the living world.
Article first published 17/10/12
Comments are now closed for this article
There are 4 comments on this article.
Jerome Ravetz Comment left 18th October 2012 18:06:48
"they are more likely to survive in the environment with unknown consequences for health and the ecology of our aquatic systems."
This comes in as an afterthought. So even for i-sis our ignorance is still less important than our knowledge.
Mae-Wan Ho Comment left 18th October 2012 20:08:49
Hi Jerry,
Sorry it seems like an afterthought to you. But definitely not! I see an obligation to inform on what is known as far as possible and then point out the areas of our ignorance. You would not want me to speak from a position of ignorance do you?
Regards, maewan
Linus Hollis, ScD Comment left 20th October 2012 18:06:55
Prion treatment! Obvious after saying it, true, but reversing the RNA cascade with aptamers should be studied asap! RE environmental contamination: if proven, treatment centers with biowaste controls will add costs, but not terribly high, while the life-saving possibilities are devoloped. Bravo!
Jerry Ravetz Comment left 20th November 2012 03:03:10
Sorry - I didn't want to be negative about your work, of course.
It's just that in your article the warning came at the very end. This seemed too similar to what one would expect in a mainstream publication, where 'absence of proof of harm is proof of absence of harm'. All the very best - Jerry